Patient vs Summary Mode
Source:vignettes/a07_patient_vs_summary_mode.Rmd
a07_patient_vs_summary_mode.RmdIntroduction
CohortContrast Viewer supports two data modes:
-
Patient: uses patient-level parquet files (for
example
data_patients.parquet). -
Summary: uses precomputed aggregate parquet files
(for example
concept_summaries.parquet).
In the Studies table, the Mode column
shows this in one word: Patient or
Summary.
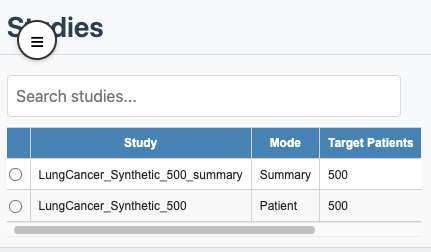
Study selection with mode column
How each mode is produced
-
Patient mode: produced directly by
CohortContrast(..., createOutputFiles = TRUE). -
Summary mode: produced by
precomputeSummary(studyPath = ..., outputPath = ...).
The bundled example studies include one patient-mode study and one summary-mode study:
patientStudyPath <- system.file("example", "st", "lc500", package = "CohortContrast")
summaryStudyPath <- system.file("example", "st", "lc500s", package = "CohortContrast")
data.frame(
study = c("lc500", "lc500s"),
mode = c(
CohortContrast::checkDataMode(patientStudyPath)$mode,
CohortContrast::checkDataMode(summaryStudyPath)$mode
)
)
#> study mode
#> 1 lc500 patient
#> 2 lc500s summaryThis mirrors the distinction shown in the Viewer study-selection table.
summary_result <- CohortContrast::precomputeSummary(
studyPath = file.path(getwd(), "studies", "LungCancer_1Y"),
outputPath = file.path(getwd(), "studies", "LungCancer_1Y_summary"),
clusterKValues = c(2, 3, 4, 5),
minCellCount = 5
)
# Open viewer and load the summary study
CohortContrast::runCohortContrastViewer(
dataDir = file.path(getwd(), "studies")
)What is different in the UI
-
Mappings merge actions:
- Patient: available (
Manual Merge,Hierarchy Suggestions,Correlation Suggestions). - Summary: disabled (history is still visible).
- Patient: available (
-
Clustering updates:
- Patient:
Reclusterruns live clustering. - Summary: clustering uses precomputed artefacts for selected
kwhen filters are applied.
- Patient:
-
Data granularity:
- Patient: row-level patient events are available.
- Summary: only aggregate summaries are available.